Particle tracking velocimetry for CFD validation
When we simulate a system, how do we know the model is correct — especially when we can't see inside a typical stainless steel bioreactor?
The problem
CFD simulations of bioreactor internals depend on input parameters that are rarely measured directly — they're guessed or borrowed from literature for different geometries. Without ground-truth flow data, there's no way to close the loop between simulation and reality.
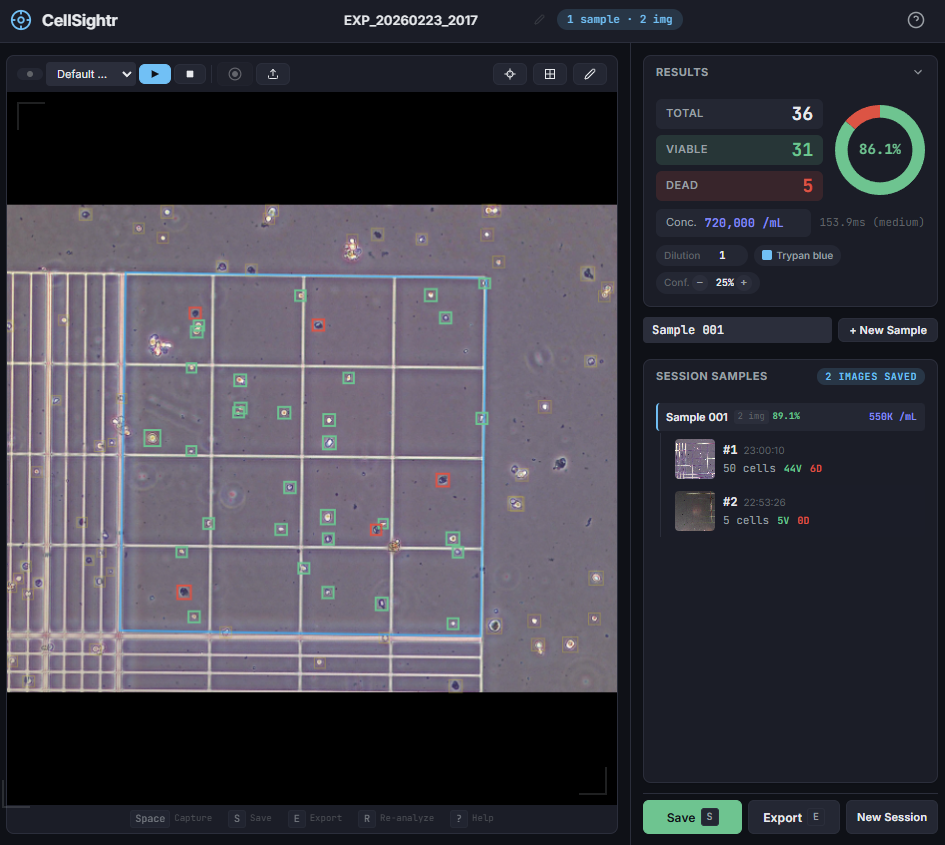
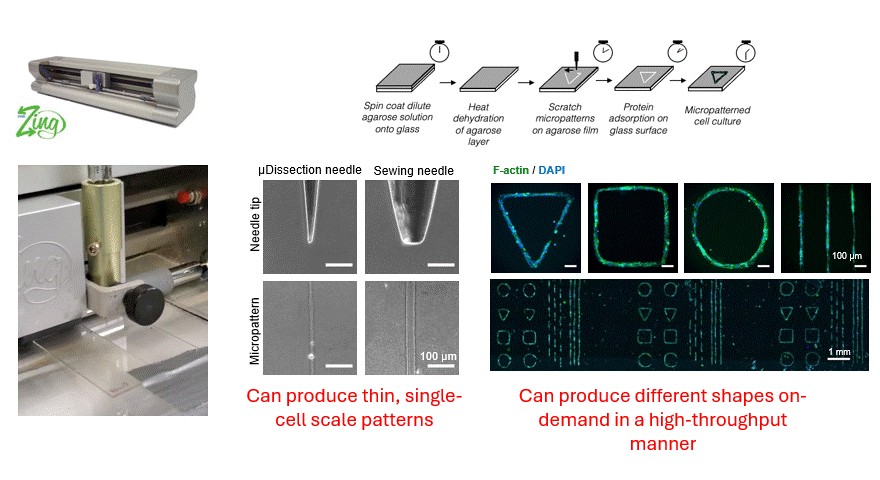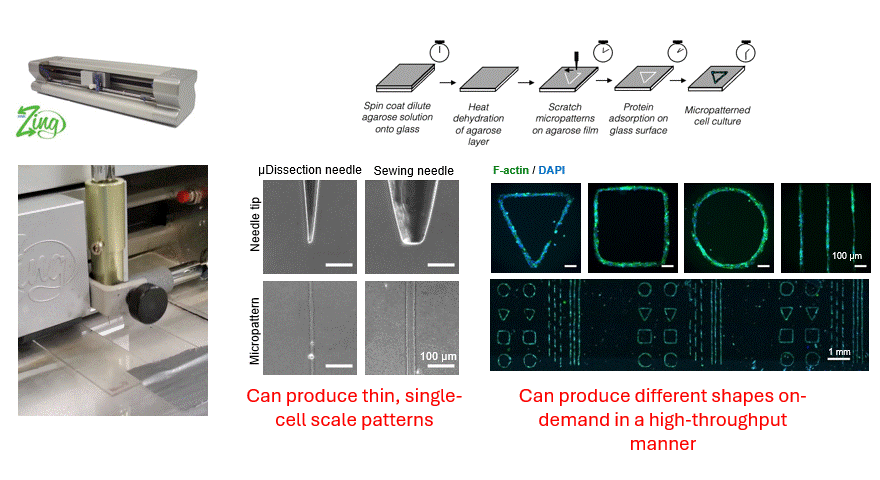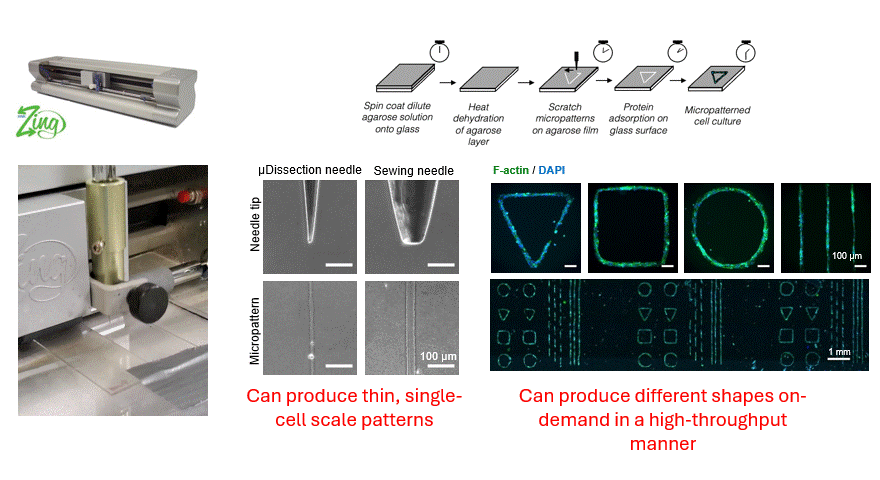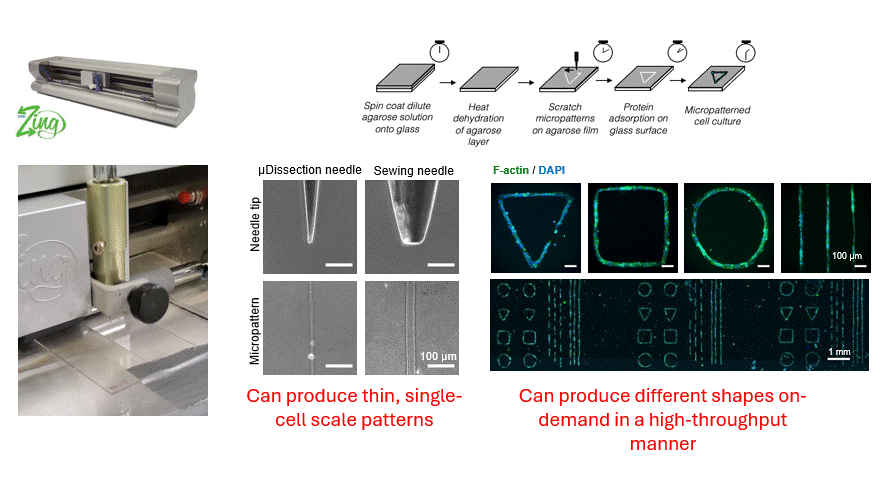
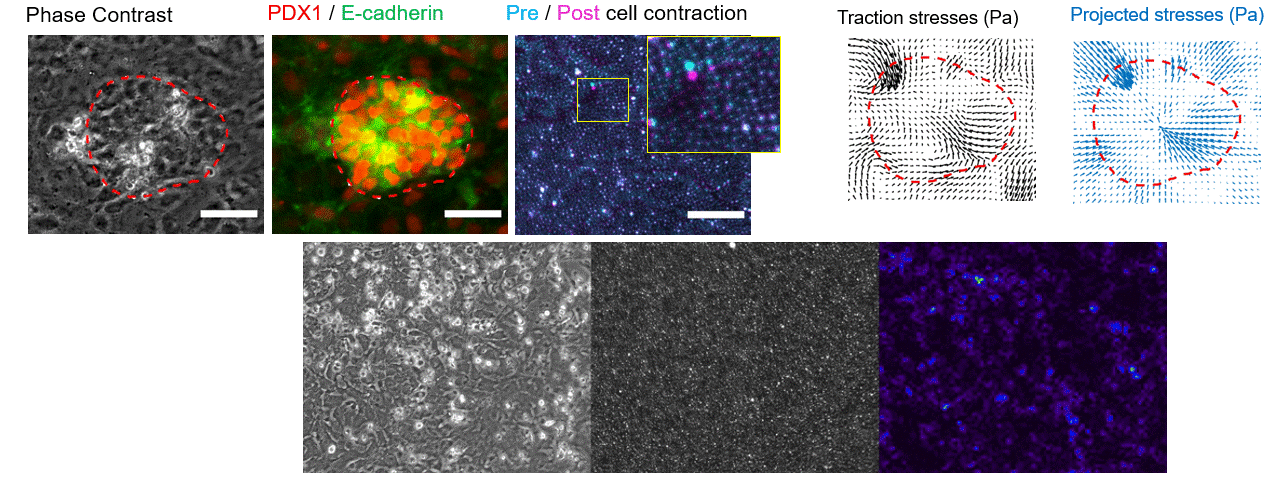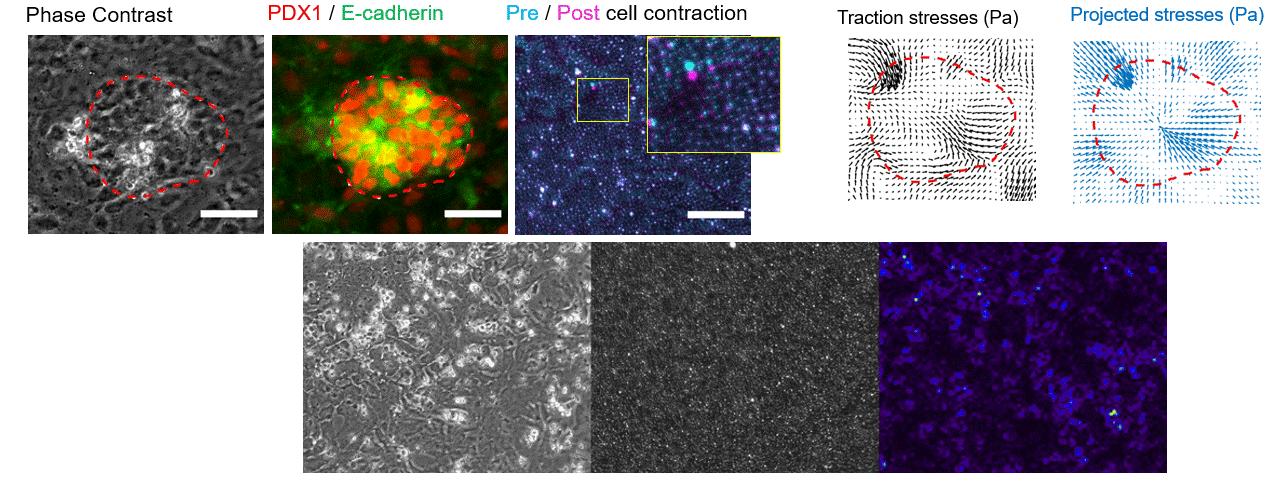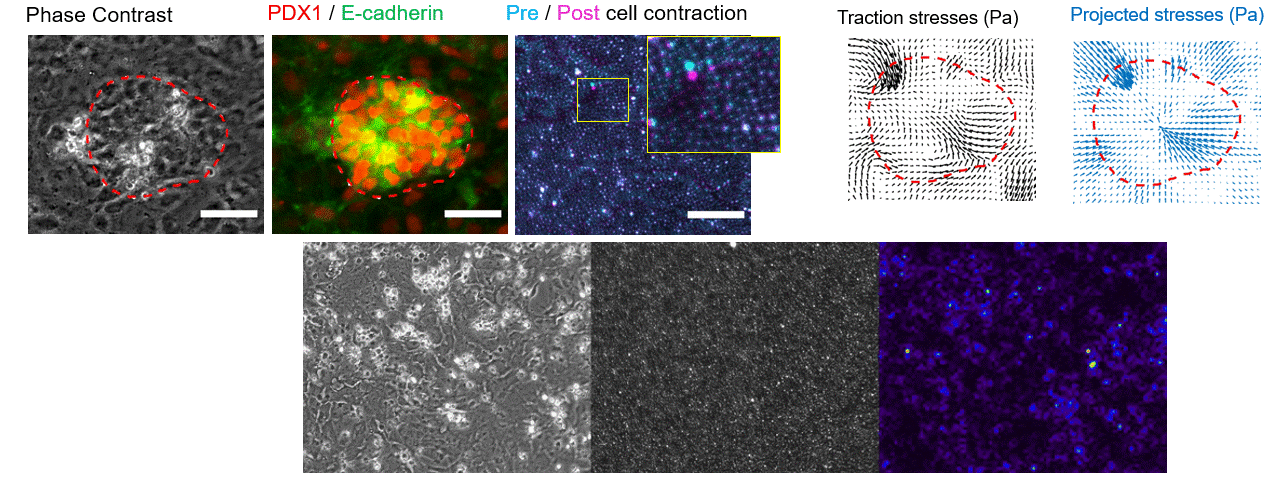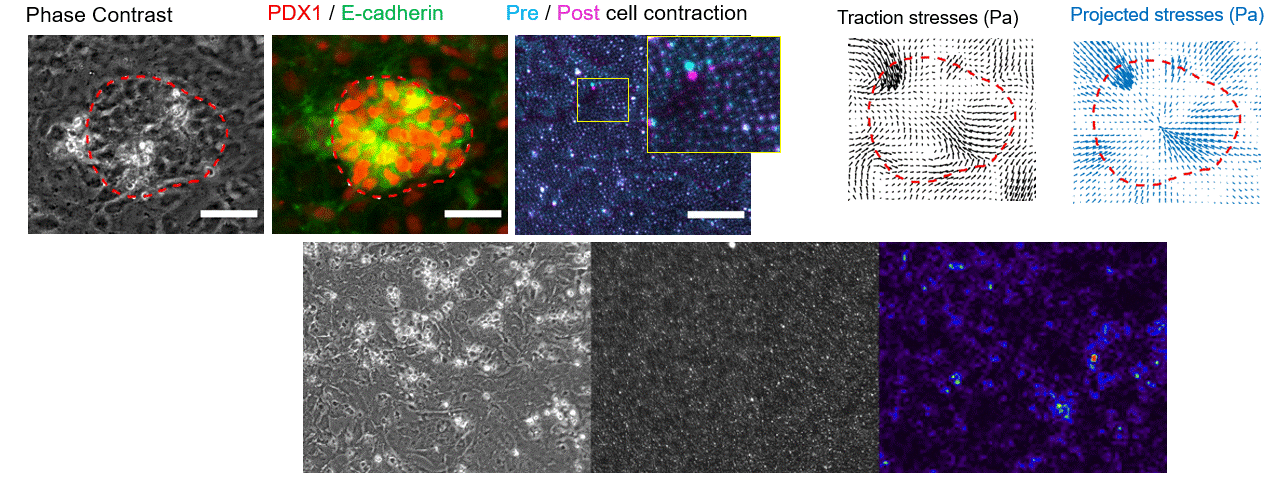
Making flow visible
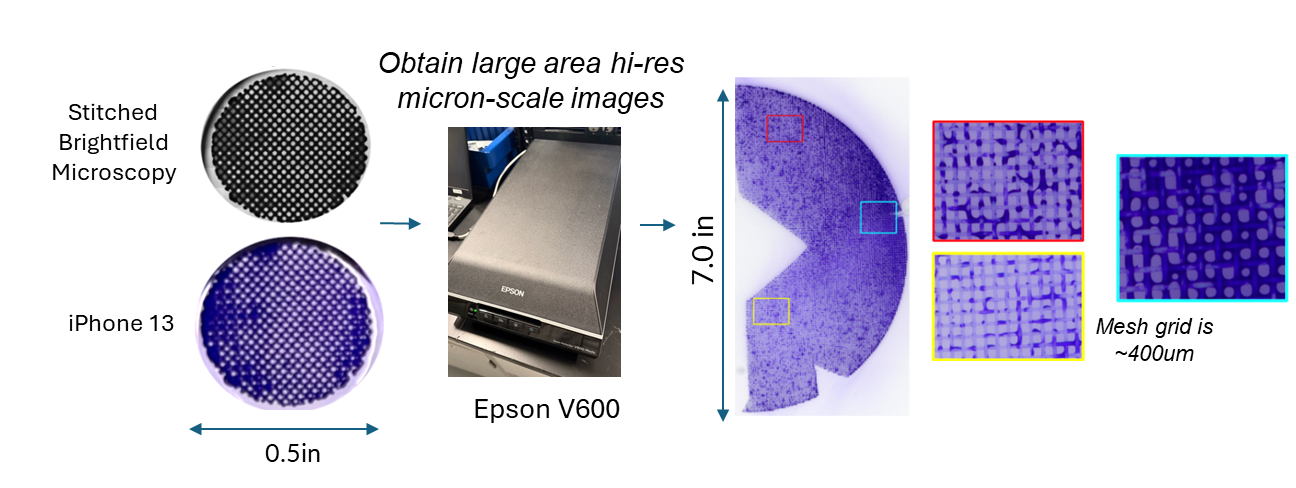
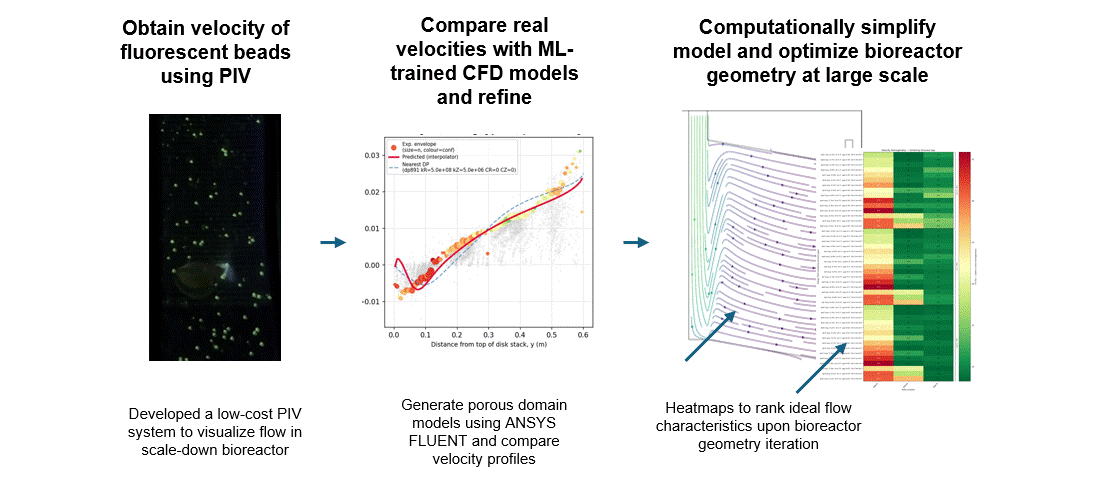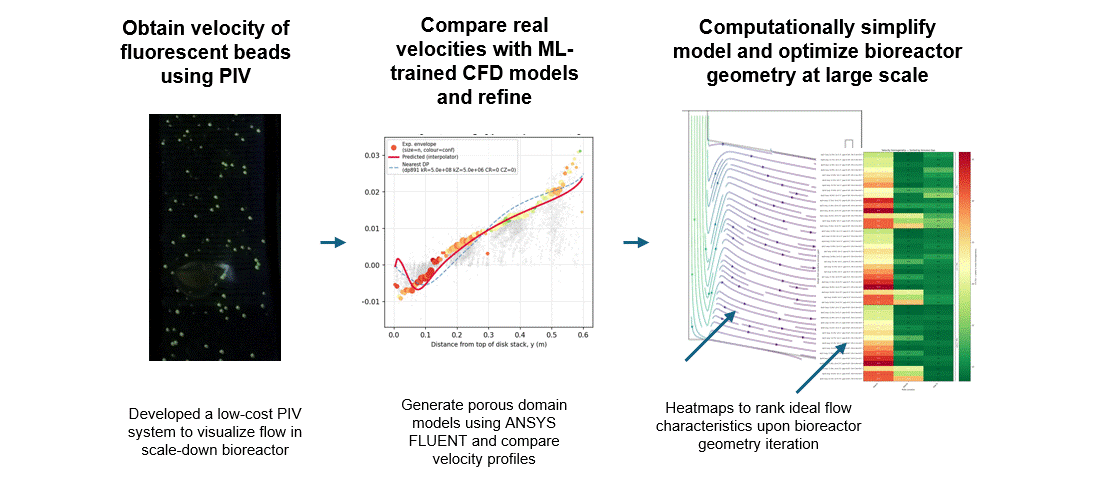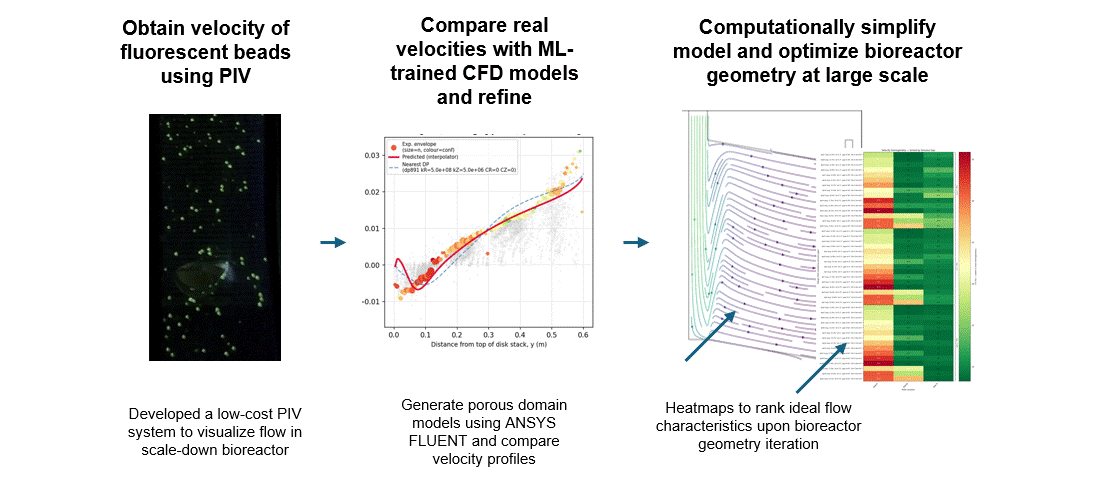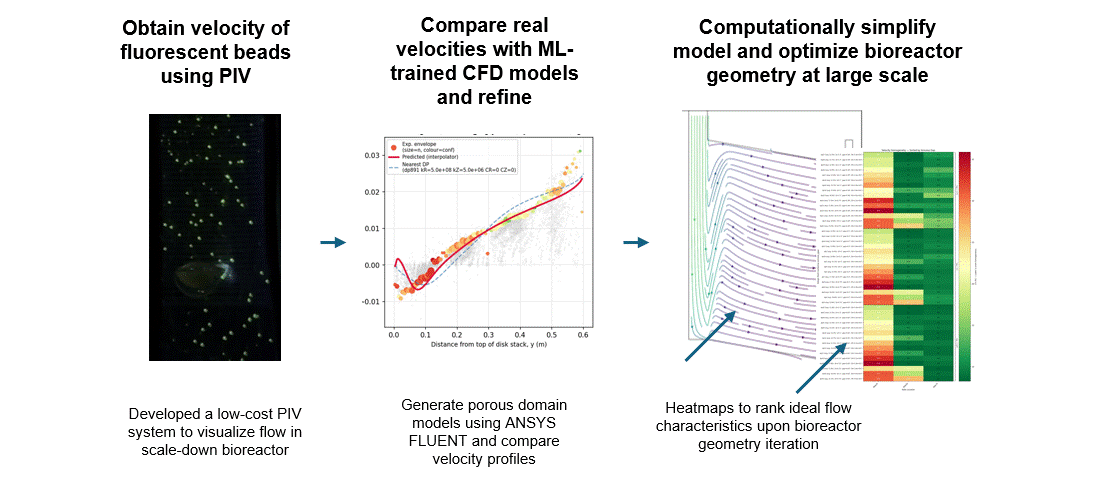
To get that ground truth, I led a team to design fully transparent bioreactor analogs that allow optical access to the flow field. We seeded the fluid with fluorescent tracer beads, illuminated a single plane, and captured footage using consumer phone cameras — no expensive laser-based PIV rigs required. From there, we built a video and image analysis pipeline to extract velocity fields via particle tracking velocimetry (PTV).
Inverse modeling
A surrogate model trained on CFD simulation ensembles across a wide parameter space maps experimental velocity profiles to the simulation input parameters that best reproduce the observed flow. These calibrated parameters are then used to run the final validated simulation.
Impact
With a validated model in hand, we could iterate on our novel bioreactor design computationally rather than solely through expensive and time-consuming physical prototypes. This gave us a predictive tool to evaluate design changes at larger scales before committing to fabrication — significantly accelerating the design-build-test cycle for scale-up.
Note: images and video shown here are representative demonstrations, not the actual bioreactor system.